|
Our BeanBeetleMicrobiome app Tutorial provides additional guidance. The app allows students to identify core and unique taxa, view rarefaction curves, visualize taxonomic diversity, and explore alpha and beta diversity. To conduct community analysis of the level-5 (family-level) taxonomy data that are generated by the Purple line on DNA Subway, we developed the BeanBeetleMicrobiome app ( ). Use our DNA Subway Tutorial and DNA Subway Video Tutorial for guidance on how to use the Purple line. The Purple line is a web-based implementation of the QIIME2 pipeline. Analysis of MiSeq Dataįor bioinformatic analysis of MiSeq data, we suggest using Purple line in DNA Subway ( ). html) or with the help of software such as snapgene or snapgene-viewer (.gbk). fa, and you will need to change the command accordingly. quality sequencing of your whole plasmid(s) at an affordable price (25 CAD. The extension for the sequence file might be. Then, type “cat *.fasta > combined.fasta”. To combine fasta files on a Mac, navigate to the folder with the files in the Terminal. For directions on how to combine fasta files (which are text files) in Windows, see. The sequences for each individual colony can be submitted separately or the fasta files for each colony can be combined into a single fasta file first. More accurate identification is possible using a curated database of 16s rRNA sequences, such as Silva ( ). For more details, see our Student handout on BLAST analysis of sequencing data. Sequences can be trimmed in SnapGene Viewer and then exported as fasta files.īacterial species (or more typically genera) can be identified using a BLAST search on Genbank. An alternative is to have students view the chromatograms and quality data in a viewer such as SnapGene Viewer ( ). Sanger sequencing of colony-based PCR products will result in chromatogram files and sequence (or fasta) files.Ĭhromatogram files in standard PDF format can be viewed and fasta files trimmed in a text editor. You can access the Features panel by clicking the flag icon in the right-side menu.Analysis of colony-based PCR 16s rRNA data 
Genetic codes on translations import as /transl_table propertiesĬustom fields, where present, are captured, viewed, and editable in the Features panel. This enables users to import, store, edit, and export all data associated with GenBank features, while modernizing the experience of creating, editing, and managing annotations and translations in Benchling.įeatures of any other type create annotations In Benchling, these custom fields are user-defined name/value pairs. Highest quality HD recorded MP3 downloads. Some properties on GenBank features are standard, like /notes, /type, and /label, but GenBank allows users to encode their own properties to store user- or customer-specific data. SnapGene Viewer is free software that allows molecular biologists to create. Tip: If you an encounter an import error, ensure your file extensions are formatted correctly. Importing other file formats sometimes work, but users should depend on them working with caution. Tip: If you don't have a sequence to import but want to practice uploading sequence files, download pBR322 to your computer. 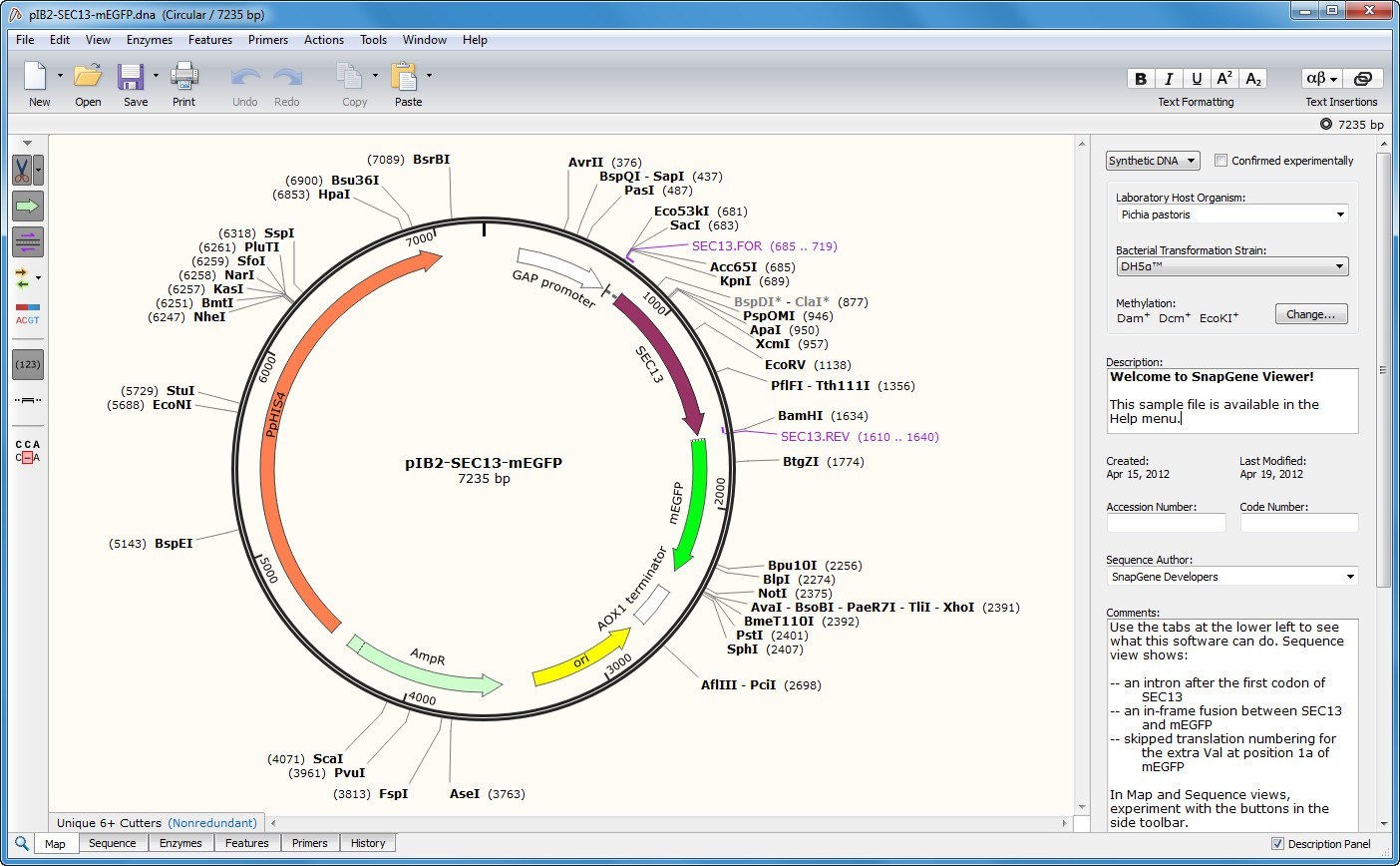
(Optional) In the menu that displays after uploading the file, update the file location, sequence topology, or tags associated with the file, as needed.Ĭlick Open Sequence to open the file, or click Close to close the modal. Select the folder to save your sequence to.Ĭlick Choose a file and select the file(s) from your computer, or drag and drop the file(s) into the box. To import sequences saved to your computer: Import a sequence file from your computer Note: The columns of the table are defined in the Registry configuration.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
